Research Article
Akanksha Mehra
Akanksha Mehra
Department of Plant Molecular Biology and Biotechnology, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: akankshamehra1999@gmil.com
Taruna Borule
Taruna Borule
Department of Plant Molecular Biology and Biotechnology, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: tarunaborule94@gmail.com
Vipin Kumar Pandey
Vipin Kumar Pandey
Department of Genetics and Plant Breeding, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: vipinpandey102@gmail.com
Shivani Singh Rana
Shivani Singh Rana
Department of Plant Molecular Biology and Biotechnology, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: shivanisinghrana885@gmail.com
Pradnya Raut
Pradnya Raut
Department of Plant Molecular Biology and Biotechnology, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: pradnyaraut628@gmail.com
Ravi Ratan Saxena
Ravi Ratan Saxena
Department of Agricultural Statistics and, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: drravisaxena@gmail.com
Satish Balkrishna Verulkar
Satish Balkrishna Verulkar
Department of Plant Molecular Biology and Biotechnology, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: satishverulkar@gmail.com
Vikas Kumar Singh#
Vikas Kumar Singh#
International
Rice Research Institute, Los Banos, 4031, Philippines.
# IRRI South Asia Hub, Patencheru Hyderabad 502324, India
E-mail: v.k.singh@irri.org
Uma Maheshwar Singh@
Uma Maheshwar Singh@
International
Rice Research Institute, Los Banos, 4031, Philippines.
@ IRRI South Asia Regional Centre, Varanasi, Uttar Pradesh 221106, India.
E-mail: uma.singh@irri.org
Arvind Kumar@
Arvind Kumar@
International
Crops Research Institute for the Semi-Arid Tropics, Patancheru, Telangana
502324, India.
@IRRI South Asia Regional Centre, Varanasi, Uttar Pradesh 221106, India.
E-mail: Arvind.Kumar@icrisat.org
Shubha Banerjee*
Shubha Banerjee*
Corresponding author
Department of Plant Molecular Biology and Biotechnology, Indira Gandhi Krishi Vishwavidyalaya, College of Agriculture, CoA, Raipur, Chhattisgarh, India.
E-mail: shubha.banerjee@igkv.ac.in, shubha19.banerjee@gmail.com.
Abstract
The
Glycemic Index (GI) is an important quality trait of rice that determines
consumer preference. GI is dependent on the relative proportion of
amylose/amylopectin, soluble and resistant starch which in turn affects the
ease of hydrolysis of the starch. To understand the relation between amylose
content (AC) and GI the activity of starch synthase (SS) and branching enzyme
(BE) was studied in the developing grain at the milking stage along with the
expression analysis of 9 genes encoding enzymes involved in amylose synthesis,
amylopectin synthesis and sucrose transport in flag leaf tissue. Expression
analysis was done in 6 genotypes with high and low GI values viz., Ashmber-GI
88.4, Makro- GI 67.3, IR 64- GI 63.4, Madhuraj 55- GI 62.7, Noh hai- GI 82.9
and Swarna- GI 69.2. Out of the 6 genotypes the activity of SS enzyme, related to
amylopectin synthesis, was found highest in low GI rice genotypes (Makro,
Madhuraj 55 and Swarna) and highest in Ashmber and Noh hai (high GI genotypes).
The activity of BE was recorded maximum in Makro, Ashmber and IR 64 and minimum
in Swarna, establishing the role of SS enzymes in the formation of soluble
starch. The expression profile indicated
that the amylose and total starch (TS) content are not determined by a major
gene but are affected by the interaction of sucrose transporters and synthase
enzymes that in turn determined amylose, amylopectin, and TS concentration. The gene expression and enzymatic activity showed
interactions in the metabolic regulation of starch biosynthesis, transport, and
packaging grains, which significantly influenced the glycaemic index.
Abstract Keywords
Starch
biosynthesis, glycemic index, amylose content, sucrose transporter genes, rice.
GI is an important quality trait of
rice. It is determined by the relative amount of amylose, amylopectin, and the proportion
of soluble and resistant starch. Glucose is stored as starch in rice grains.
The essentially linear -polyglucan amylose and the branched -polyglucan
amylopectin are the two main components of starch. Amylose is a linear polymer
made up of glucan chains that are 1,4-linked and contain a very small number of
branches joined by 1,6-glycosidic linkages. While the amylopectin molecule is
larger than the amylose molecule and contains abundant α-1,6-branches that
connect α-1,4-linked glucan chains that make up a structural framework of
repeated amorphous and crystalline lamellae.
The granule-bound starch synthase
(GBSS) and soluble starch synthase (SS) enzymes catalyse the reactions for the
elongation of 1,4-chains of amylose and amylopectin, respectively. Amylopectin
is made up of tandem-linked clusters (roughly 9–10 nm in length each), where
linear α-1,4-glucan chains are regularly branched via α-1,6-glucosidic linkages. The
crystalline organisation of the starch granule is facilitated by the unique
structure of amylopectin [1-6]. The rate of
synthesis of amylose and amylopectin polymer, their organization in rice grains
and packaging into granules determine the overall GI of a variety. Varietal
differences and environmental factors are known to influence the fine structure
of starch clusters resulting in changes in their functional qualities [7]. Earlier studies have reported the function of
the starch synthesis and sugar transport related enzymes in rice grains and
leaves. Only a few recent studies have specially observed the relationship of
starch synthesis and transport with the rate of grain filling in rice [8]. Varietal differences in the packaging of
starch in rice grain and grain density as well as compactness have been
observed. Although the change in temperature during grain maturity is known to
affect the packaging of photosynthates in rice grains [9],
cooking quality, AC, and other quality related aspects [10],
but none of the studies have reported a direct effect on GI value. These
observations indicate the possibility of correlation of these factors with the glycaemic
index (GI) value of rice thus the study of genes and enzymes related to starch
biosynthesis and sugar transport was conducted in Indica rice varieties with
different maturity periods and contrasting GI values.
ADP-glucose pyrophosphorylase (AGPase),
starch synthase (SS), starch branching enzyme (SBE), and starch de-branching
enzyme (DBE) are the key enzymes involved in starch biosynthesis. These enzymes
are highly regulated and the limiting reaction is catalysed by AGPase, which is
the first enzyme in the starch biosynthesis pathway. It occurs in amyloplasts
by converting glucose 1-phosphate and ATP to ADP-Glucose and inorganic
pyrophosphate (PPi). SS can be further classified into soluble starch synthase
(SSS), which mostly produces amylopectin, and granule-bound starch synthase
(GBSS), which produces amylose and the extra-long-chain part of amylopectin [11]. SBEs are members of α- amylase family. The branching enzyme
introduces a branching in the polyglucan's α-1, 4-glucan chain and then reattaching
the cleaved chain to an acceptor chain via an α-1,6-glucan linkage, creating a branch
in the same or another chain [12,13].
The starch so synthesised is
transported to sink tissue through sucrose transporters. There are five types
of SUTs known in rice. SUT, a class of sucrose-proton symporters found only in
plants, is crucial for sucrose acquisition from the intercellular apoplast
outside of the cell into the cytosol. Moreover, sucrose must first be uploaded
into the phloem before it can be transported across vast distances via the
symplastic pathway. Additionally, a tiny amount of sucrose may leak out in the
apoplast during symplastic transport within the phloem; bringing this sucrose
back into the phloem also requires the involvement of the SUT transporter [14]. The apoplastic mode of transport majorly
contributes to the transport in developing rice grain i.e., sink tissue. The
sucrose is uploaded into the phloem from the sheath tissue of flag leaves which
serves as source tissue. Between source and sink tissues, there are different
metabolic pathways involved in starch production. The carbon in starch is
derived from fructose-6-P in the Calvin-Benson cycle in photosynthetic tissues,
whereas it is derived from sucrose in sink tissues, which is transported from
source tissues through the phloem [15].
Sucrose is the primary form in which assimilate, produced from photosynthetic
source tissues like the flag leaf blade, is transferred to sink tissues via the
long-distance vascular pathway. Grain that accumulates starch in the endosperm
is the main sink tissue in cereals. Prior to heading, other tissues, such as
the leaf sheath and stem, may also serve as temporary sink tissues. A
continuous, mature transport phloem network carries the sucrose to these
different sink tissues [16].
The granule-bound starch synthase I
(GBSS/Wx) is vital for amylose synthesis while SS I and II play a role in
extending the amylopectin chain [17,12].
Rice genotypes with differences in GI values have shown allelic variation at
Waxy locus [18]. Genetic variation in the
waxy locus leads to difference in grain transparency and cooking quality in
rice, the rare alleles of the waxy locus have been identified that govern AC
and eating as well as cooking quality of japonica rice [19,
20]. The BEs are known to introduce α-1,6-linkage via and activity of
DBEs leads to formation of the crystalline amylopectin by removal of
a-1,6-linkages [21,17]. Since the starch
biosynthesis and transport in rice is largely governed by genes such as glucose
phosphorylase (AGPLs), starch synthase (SSs), branching enzyme (BEs),
de-branching enzyme (DBE), and sucrose transporters (SUTs) in developing seeds
and leaves, [22, 5] therefore the expression
of nine genes viz. OsAGPL1, OsAGPS1, OsSS2a, OsGBSS2, OsWx, OsBE-2a, OsSUT-1,OsSUT4,
OsSUT- 5 were analysed in the flag leaf tissue. The assay of the corresponding
enzymes such as starch synthase and branching enzyme was also done in
developing endosperm. The pattern of gene expression of the nine genes was
analysed for correlation with grain AC and Glycaemic index value of the grains
harvested from the same plant.
2. Materials and methods
2.1 Plant material
The experimental material for
the present study consisted of 208 diverse rice genotypes which were evaluated
in an augmented block design at the field experimentation centre of the
Department of Plant molecular biology and biotechnology, Raipur College of Agricultural,
Chhattisgarh during the Kharif season (June- December), 2021. The recommended
cultural practices were followed to raise the crop.
2.2 Analysis of AC, predicted
glycemic index and grain quality traits
The observations were recorded
for quality characters viz., grain length, grain width, length/breadth (l/b)
ratio, chalkiness, milling percentage, Head Rice Recovery (HRR) percent and
agronomic traits viz., plant height, no. of panicle/plant, panicle length and
panicle weight as per the SIS of rice by IRRI (2002).
ANOVA of all the above traits was calculated using an augmented design layout
and Pearson correlation coefficient was also calculated by using R software.
Biochemical analysis of each genotype for estimation of AC (as per the method
by [23] and glycemic index [24] was carried out. Based on the AC (high and low) and glycemic
index (high and low) six genotypes were selected for enzyme assay and gene
expression analysis. Seeds of the 6 rice varieties, Ashmber (GI- 88.44), Makro (GI- 67.37), Madhuraj-55 (GI-
62.79), IR64 (GI- 63.46), Swarna
(GI- 69.27) and Noh Hai (GI- 82.85), were planted in pots. Seedlings were
transplanted to the field 21 days after sowing (DAS) and tissues were harvested
in liquid nitrogen at the milking stage (usually 7 to 10 days
after heading) for enzyme assay. Flag leaf tissue was used for
gene expression analysis since it is considered to contribute maximum to the
developing grains. All leaf samples were instantly frozen in liquid nitrogen
after being cut off from the plant and kept for further investigation.
2.3 Enzyme assay
Assay activity of Starch
synthase (SS) and Branching enzyme (BE) was carried out at the milking stage
(10 days after pollination) in endosperm tissue. The assay of Starch synthase
and Branching enzyme was done as per the method followed by [25]. For assay, 25 hulled grains were used that
were harvested at the milking stage. The embryo and pericarp were removed, and
the grains were homogenised with 5 ml of the buffer solution in an ice-cold
mortar along with 12.5% (v/v) glycerol and 5% (w/v) insoluble
polyvinylpyrrolidone 40. Unless otherwise noted, the homogenate was centrifuged
at 10,000 x g for 5 min, and the resultant supernatant was utilised to prepare
the enzymes [25]. Starch synthase
-The analysis was carried out in a reaction mixture of 280 μl with 50 mM HEPES-NaOH (pH
7.4), 1.6 mM ADP glucose, 0.7 mg amylopectin, 15 mM DTT, and enzyme
preparation. The enzyme was inactivated by putting the mixture in a boiling
water bath for 30 seconds twenty minutes after the reaction began. The mixture
was then treated with 100 μl of a solution containing pyruvate kinase (1.2 unit), 200 mM KC1, 10 mM
MgCl2, 4 mM PEP, and 50 mM HEPES-NaOH (pH 7.4), and incubated for 30
min at 30°C. After the reaction by starch synthase the ADP is turned into ATP and
the resultant solution was heated in a boiling water bath for 30 seconds before
being centrifuged at 10,000xg for five minutes.A mixture of the supernatant
(300 μl), 50 mM HEPES-NaOH (pH 7.4), 10 mM glucose, 20 mM MgCl2, and
2 mM NADP was added. The increase in absorbance at 340 nm following the
addition of 1 /A of hexokinase (1.4 unit) and G6P dehydrogenase (0.35 unit)
each served as a marker for the enzymatic activity [25].
Branching enzyme- The assay was carried out in a reaction mixture
of 200 μl with 50 mM HEPES-NaOH (pH 7.4), 5 mM G1P, 1.25 mM AMP, phosphorylase A (54
unit), and enzyme preparation. By adding 50 μl of 1 N HC1, the reaction was
stopped. 700 μl of 0.1% I2, 0.1% KI, and 500 μl of dimethyl sulfoxide were added to the solution.
At 540 nm, the enzyme activity was measured using a spectrophotometer. The
amount that causes a rise in absorbance of one unit at 540 nm in one minute was
used to define one unit of enzymatic activity [25].
2.4 RNA extraction and cDNA preparation
The
flag leaf of 55 days old rice genotypes was collected for RNA extraction.
Samples were crushed using liquid nitrogen and total RNA was extracted
immediately using the Trizol reagent method [26].
Using a Nanodrop spectrophotometer (ND 1000), the amount of RNA is measured at
260 nm. To measure absorbance, 1 µl of isolated RNA was put over the
nanodrop's tip. For each sample, the absorbance ratios (A260/A280) were
recorded in order to assess the samples and determine the RNA purity. For pure
RNA, an acceptable absorbance ratio (A260/A280) was between 1.9 and 2.1. The
extracted RNA was also loaded on 0.8% gel to assess the integrity of the RNA. The RNA was then used for cDNA synthesis using Bio-Rad
iScript cDNA Synthesis Kit as per the manufacturer’s instructions.
2.5 Semi quantitative RT- PCR
The expression of starch related genes
was investigated using semi-quantitative RT-PCR. The cDNAs synthesized was used
for the expression analysis using gene specific primers (Table 1). The
resultant PCR products were resolved on 1.5 % Agarose gel at 70V. The presences
of amplicons with their respective intensity were recorded under gel
documentation. The relative intensity of amplicons provided basis for the quantification
of the level of expression of the gene as high, moderate, low and negligible.
Table 1. Target genes for analysis of expression profile
|
Traits |
Mean |
S.D. |
S.E. |
Range |
Skew-ness |
Critical difference (5%) |
Coefficient of Variance |
Mean sq. (un- adjusted) |
Mean sq. (adjusted) |
|
GL |
6.02 |
0.71 |
0.05 |
4.01-8.81 |
0.24ns |
0.49 |
3.46 |
0.19** |
0.57** |
|
GW |
2.26 |
0.24 |
0.02 |
1.58-3.03 |
0.17ns |
0.21 |
3.93 |
0.03* |
0.07** |
|
l/b
ratio |
2.69 |
0.45 |
0.03 |
1.57-3.97 |
0.49** |
0.38 |
5.83 |
0.06ns |
0.23** |
|
Chalki |
6.71 |
2.43 |
0.18 |
1.0-9.0 |
-0.93** |
3.1 |
19.75 |
0.9ns |
5.55** |
|
Milling |
64.59 |
9.57 |
0.72 |
29.46-85.91 |
-0.85** |
17.39 |
11.35 |
223.47** |
94.66* |
|
HRR |
52.84 |
12.72 |
0.96 |
12.6-79.7 |
-0.55** |
15.32 |
12.3 |
212.71** |
157.75** |
|
PH |
147.99 |
25.87 |
1.95 |
83.96-198.21 |
-0.41* |
38.65 |
11.23 |
1642.26** |
861.89** |
|
P/p |
7.23 |
1.55 |
0.12 |
4.11-12.71 |
1.04** |
2.91 |
16.86 |
2.61ns |
2.51ns |
|
Pl |
25.83 |
2.47 |
0.19 |
18.02-32.6 |
-0.16ns |
5.17 |
8.43 |
2.89ns |
5.97ns |
|
PW |
13.08 |
3.88 |
0.29 |
3.86-25.65 |
0.41* |
6.25 |
20.4 |
16ns |
17.4ns |
|
DTM |
138.96 |
10.97 |
0.83 |
115.4-164.03 |
0.01ns |
17.87 |
5.38 |
140.56ns |
133.83** |
|
Note: 1. ns P > 0.05; * P <= 0.05; ** P <= 0.01; 2. GL= Grain length, GW= Grain width, l/b= Length/Breadth ratio, Chalki= Chalkiness, Milling= Milling percent, HRR= Head rice recovery, PH= Plant height, P/p= No. of panicles/plant, PL= Panicle length, PW= Panicle weight, DTM= Days to maturity, S.D.= Standard deviation, S.E.= Standard error. | |||||||||
3. Results and discussion
3.1 Analysis of grain quality traits:
Variation of all the traits related to
grain quality and agronomic performance was studied (Fig. 1 and 2) and their
range was recorded (Table 2).
Figure 1. Frequency distribution of Grain
quality parameters viz., Fig. 1a- Grain length (mm), Fig.1b-Grainwidth(mm),
Fig.1c-Length/Breadthratio, Fig.1d-Chalkiness, Fig. 1e- Milling percent,
Fig.1f- Head rice recovery in a diverse rice genotype.
Figure 2. Frequency distribution of
Agronomic traits viz., Fig.2a- Plant height (cm), Fig.2b- No. of
panicles/plant, Fig. 2c- Panicle length (cm), Fig. 2d- Panicle weight (g), Fig.
2e- Days to maturity.
For analysis of variance, augmented block design was applied in the diverse rice genotypes. Observing the grain quality traits and agronomic traits shows significant variation in traits viz., grain length, grain width, length/breadth ratio, chalkiness, milling percent, head rice recovery (HRR), plant height, no. of panicles per plant, panicle length and days to maturity (Table 2).
The range of grain length and width varied significantly (Table 2) with Altamira-9 genotype having maximum grain length and Kuneng genotype having maximum grain width. Similarly, minimum grain length and grain width values were recorded in improved Sambha mahsuri and Swarna respectively. From grain length and width l/b ratio (length/breadth ratio) is calculated. The genotypes showing maximum and minimum l/b ratio are RNR 15048 and Thavalkannan.
Table 2. Analysis of variance of phenotypic traits of diverse rice genotypes viz., grain length, grain width, l/b ratio, chalkiness, milling percentage, HRR percent and agronomic parameters viz., plant height, panicle length, no. of panicle per plant, panicle weight with AC, TS and predicted glycemic index.
Enzyme | Gene | Expected amplicon size (bp) | Chr. | Primer sequence | Ref. |
ADP-glucose pyrophosphorylase large subunit | OsAGPL1 | 140 | 5 | f-CATCAAGGACGGGAAGGTCA r-ACTTCACTCGGGGCAGCTTA | [44] |
ADP-glucose pyrophosphorylase small subunit | OsAGPS1 | 97 | 9 | f-AGAATGCTCGTATTGGAGAAAATG r-GGCAGCATGGAATAAACCAC | [44] |
Starch synthase IIa | OsSSIIa | 54 | 6 | f-GGCCAAGTACCAATGGTAAA r-GCATGATGCATCTGAAACAAAG | [38] |
Granule-bound starch synthase II | OsGBSSII
| 246 | 7 | f-AGGCATCGAGGGTGAGGAG r-CCATCTGGCCCACATCTCTA | [38] |
Granule-bound starch synthase I | Wx6 | 852
| 6 | f-CTCTTATTCAGATCGATCAC r-CCATATGTGCTTATAAGGAC | [38] |
Starch branching enzyme IIa | OsBEIIa | 128 | 4 | f-GCCAATGCCAGGAAGATGA r-GCGCAACATAGGATGGGTTT | [45] |
Sucrose transporter 1 | OsSUT1 | 125 | 3 | f-AGTTCTGGTCGGTCAGCAT r-ACCGAGGTGGCAACAAAG | [46] |
Sucrose transporter 4 | OSSUT4 | 59 | 2 | f-TTTGGCTGAGCAGAACACCA r-ATGTCATTCGGGCAGAGCTT | [46] |
Sucrose transporter 5
| OsSUT5 | 64 | 2 | f-CTAGTGCGAAACTCCATCAAA r-AAAATATTTGGGTTTCCTGAGAT | [46] |
Chalkiness is an important index to assess the appearance quality of rice. Chalkiness also plays an important role in determining the processing, cooking and eating qualities of rice. It needs more attention to understand and select for the chalkiness of grains because rice with high chalkiness loose grain hardiness and becomes soft and easy to break during milling. It leads to the reduced grain milled rice rate and head milled rice rate [27]. Minimum chalkiness that is 1(absent) was recorded in 16 genotypes like- Aus 295, Gi tah and RD 15 while maximum chalkiness value 9 (large) was observed in 65 genotypes like Improved sambhamahsuri, Pulutselemyum and Dhane burwa. Milling efficiency and head rice recovery (HRR) determines the final whole-grain yield and the milled rice's broken kernel rate, which concerns the milling industry, consumers, and farmers [28]. In our study, minimum milling% was recorded in IR 4432 followed by Pa woon and Maung nyo and maximum milling% was observed in Lalat followed by Muttu samba and BW 295-5 respectively. The minimum head rice recovery was recorded in War 72-2-1-1 followed by Aus 219 and Thavalkannan and maximum head rice recovery was observed in Swarna followed by Muttu samba and Cauvery respectively. Grain length, length/breadth ratio, chalkiness, milling percent and head rice recovery are considered to be important traits that determine commercial acceptability and consumer preference of a rice variety [29, 30]. From the above findings, it may be concluded a variety having a long slender structure with higher head rice recovery and low chalkiness is desirable. Although, none of the genotype was identified which had all of the favoured grain quality traits but B 6136 E-3-TB-0-1-5 genotype possessed having long slender structure, no chalkiness with intermediate head rice recovery and moderate GI value.
In order to assess the agronomic traits of these genotypes’ traits such as plant height, date of flowering, days to maturity, number of panicles per plant, panicle length, and grain yield were also observed. All these traits showed wide genetic variation, among the 208 genotypes evaluated and analysed in the augmented design (Table 2). The plant height (PH) is a critical determinant of lodging resistance and strongly influences yield in rice, so a moderate plant height is an important basis for rice breeding [31]. Minimum plant height was recorded in Min zao 6 and maximum plant height in Eakhawngan genotypes respectively. Important agronomic characteristics associated with grain yield include panicle structure, panicle branching, spikelet production and grain morphology [32]. We recorded maximum panicle length in ARC 18502 and minimum panicle length in Cauvery. No. of panicles per plant and panicle weight also showed a high variation among the studied genotypes. Maximum and minimum no. of panicles/ plant was recorded in Qinognas and Nibari respectively while maximum and minimum panicle weight were recorded in Kam pai and Genit genotypes. The rice genotypes with desired agronomic traits such as semi dwarf plant height, higher no. of panicles per plant, panicle length and panicle weight are more suitable.
RPW 9-4 (SS1) rice variety had less plant height, intermediate panicle length and no. of panicles/plant and high panicle weight which can be commercially favourable.
Days to maturity directly influences the starch quantity and the duration of the rice genotypes is considered to play a critical role in the deposition of starch in the endosperm. Significant variations were observed in the genotypes with respect to days to maturity. WAR 72-2-1-1 was showing maximum days to maturity followed by Dudhkadar and Khao gradookchahng while Liong orn was showing minimum days to maturity (early maturing variety) followed by Chama and IR 3829.
3.2 Analysis of AC, TS and glycemic index
The frequency distribution of AC, TS and glycemic index was studied (Fig. 3 and their ranges were recorded (Table 3). For analysis of variance augmented block design was applied.
Figure 3. Frequency distribution of Biochemical traits viz., Fig. 3a-Total starch content, Fig. 3b-AC and Fig. 3c-predicted glycemic index in a diverse rice genotype.
The observations for TS and glycemic index were divided into three categories- low (<55), intermediate (56-69) and high (>70) [33]. There was significant variation observed among 208 genotypes with respect to TS and glycemic index. In our study genotypes showing maximum TS were Damnoeubkhse saut, Wanga barugulu, CT 97737 and genotypes showing maximum glycemic index were Dissi, ARC 11901 and Fu zaoxian. Similarly, minimum value for TS content was observed in Barik kudi, Safari 17 and Eloni while minimum glycemic index value was observed in, Ictacrispo 3, Ce in tsan and Eakhawngan. AC is classified into five categories- waxy (1-2%), very low (2-9%), low (10-20%), intermediate (20-25%) and high (>25%). We discovered through our research that IRRI BINI has the maximum AC, followed by Madhuraj 55 and Chama (dwarf). Similar to this, Ashmber has the minimum AC value, followed by Pulutputhi and Puttunellu. On the basis of these observations six genotypes-Ashmber and Noh hai (high glycemic index), Makro and Madhuraj 55 (high AC and low glycemic index), IR64 and Swarna (intermediate glycemic index value) were selected for enzyme assay and gene expression analysis.
Table 3. Analysis of variance of biochemical traits of diverse rice genotypes viz., AC, TS, predicted GIvalue
Traits
| Mean | S.D. | S.E. | Range | Skewness | Critical difference (5%) | CV | Mean sq. (unadjusted) | Mean sq. (adjusted) |
Amylose | 21.06 | 4.99 | 0.38 | 4.83-32.53 | -1.57 ** | 6.41 | 12.53 | 16.52 ⁿˢ | 26.63 ** |
TS | 61.99 | 5.16 | 0.39 | 50.11-84.51 | 0.82 ** | 9.69 | 7.30 | 67.08 * | 27.98 ⁿˢ |
pGI | 72.95 | 9.17 | 0.69 | 45.14-92.86 | -0.86 ** | 10.77 | 5.59 | 280.84 ** | 87.54 ** |
Note: 1. ns P > 0.05; * P <= 0.05; ** P <= 0.01; 2. Amylose= AC, TS= Total starch, pGI= predicted GI, S.D.= Standard deviation, S.E.= standard error, CV= Coefficient of variance.
3.3 Correlation of AC, TS and predicted glycemic index value with grain quality traits and agronomic traits
Analysis of correlation between the AC, TS, and glycemic index was performed with the mean values of the quality parameters and agronomic traits. Grain length shared a significant positive correlation length breadth ratio. A negative significant relationship was also reported between head rice recovery, milling percent and TS (Table 4). From this, we can conclude that there is a relationship between the strength of rice grain and starch content as the increase in TS content showed decrease in milling percentage and head rice recovery. Therefore, developing rice varieties with better head rice recovery and milling percent, the intermediate to low TS content may be used as selection criteria. Plant height also showed a significant positive correlation with panicle weight. Similarly, a significant negative correlation was also observed in no. of panicles per plant with milling percent, plant height and panicle length (Table 4).
Table 4. Correlation between quality parameters viz., grain length, grain width, l/b ratio, chalkiness, milling percentage, HRR percent and agronomic parameters viz., plant height, panicle length, no. of panicle per plant, panicle weight with AC, TS and predicted glycemic index.
| GL | GW | L/B | CHALKI | MILL | HRR | PH | P/p | PL | PW | DTM | AMY | TS | GI |
| ||||||||||||||
GL | 1 | 0.007NS | 0.681** | -0.025NS | 0.007NS | 0.021NS | -0.071NS | 0.115NS | -0.020NS | -0.018NS | -0.054NS | -0.069NS | 0.000NS | -0.136NS |
| ||||||||||||||
GW | 0.007NS | 1 | -0.712** | 0.398** | 0.105NS | 0.039NS | 0.252** | -0.050NS | 0.059NS | 0.163* | 0.005NS | -0.048NS | 0.125NS | -0.018NS |
| ||||||||||||||
L/B | 0.681** | -0.712** | 1 | -0.305** | -0.061NS | -0.002NS | -0.237** | 0.108NS | -0.059NS | -0.127NS | -0.042NS | -0.014NS | -0.095NS | -0.076NS |
| ||||||||||||||
CHALKI | -0.025NS | 0.398** | -0.305** | 1 | 0.004NS | -0.124NS | 0.203** | 0.039NS | 0.119NS | 0.146* | -0.059NS | -0.107NS | 0.083NS | 0.008NS | |||||||||||||||
MILL | 0.007NS | 0.105NS | -0.061NS | 0.004NS | 1 | 0.644** | 0.080NS | -0.146* | 0.043NS | 0.111NS | 0.021NS | -0.059NS | -0.149* | 0.116NS |
| ||||||||||||||
HRR | 0.021NS | 0.039NS | -0.002NS | -0.124NS | 0.644** | 1 | 0.101NS | -0.076NS | -0.027NS | 0.119NS | -0.025NS | -0.136* | -0.143* | 0.101NS |
| ||||||||||||||
PH | -0.071NS | 0.252** | -0.237** | 0.203** | 0.080NS | 0.101NS | 1 | -0.266** | 0.424** | 0.286** | -0.075NS | -0.178* | 0.045NS | -0.011NS |
| ||||||||||||||
P/p | 0.115NS | -0.050NS | 0.108NS | 0.039NS | -0.146* | -0.076NS | -0.266** | 1 | -0.212** | -0.126NS | 0.042NS | 0.065NS | 0.011NS | 0.002NS |
| ||||||||||||||
PL | -0.020NS | 0.059NS | -0.059NS | 0.119NS | 0.043NS | -0.027NS | 0.424** | -0.212** | 1 | 0.229** | 0.037NS | -0.097NS | 0.014NS | 0.009NS |
| ||||||||||||||
PW | -0.018NS | 0.163* | -0.127NS | 0.146* | 0.111NS | 0.119NS | 0.286** | -0.126NS | 0.229** | 1 | -0.002NS | -0.067NS | -0.019NS | 0.012NS |
| ||||||||||||||
DTM | -0.054NS | 0.005NS | -0.042NS | -0.059NS | 0.021NS | -0.025NS | -0.075NS | 0.042NS | 0.037NS | -0.002NS | 1 | 0.137* | 0.035NS | -0.053NS |
| ||||||||||||||
AMY | -0.069NS | -0.048NS | -0.014NS | -0.107NS | -0.059NS | -0.136* | -0.178* | 0.065NS | -0.097NS | -0.067NS | 0.137* | 1 | 0.152* | -0.028NS |
| ||||||||||||||
TS | 0.000NS | 0.125NS | -0.095NS | 0.083NS | -0.149* | -0.143* | 0.045NS | 0.011NS | 0.014NS | -0.019NS | 0.035NS | 0.152* | 1 | -0.074NS |
| ||||||||||||||
GI | -0.136NS | -0.018NS | -0.076NS | 0.008NS | 0.116NS | 0.101NS | -0.011NS | 0.002NS | 0.009NS | 0.012NS | -0.053NS | -0.028NS | -0.074NS | 1 |
| ||||||||||||||
Additionally, there was a significant positive correlation between TS and AC, as the AC levels signifying an increase with the increase in TS content. Plant height was also found to correlate with many traits. It showed positive significant correlation with grain weight, chalkiness as well as negative significant correlation with l/b ratio and no. of panicles/plant (Table 4). From this, we can conclude that dwarf plant varieties will have less chalkiness and slender shape as compared to tall rice varieties. A negative significant correlation of 0.178 value was also observed in plant height and AC suggesting that dwarf rice varieties have higher AC. Positive significant correlation was observed in chalkiness and grain width and panicle weight and negative significant correlation between chalkiness and l/b ratio (Table 4). Therefore, it suggests that the consumer and millers’ preference is towards long slender rice grains that have compact grain type. Days to maturity also showed a positive significant correlation with AC indicating that long duration varieties have more AC as compared to the early maturing varieties (Table 4).
According to studies conducted so far, significant negative correlation between amylose and glycemic index has been reported. However, we found non-significant negative correlation (Table 4). The non-significant findings could be due to the wide variation studied among the 208 genotypes used in this study. These genotypes belonged to various classes of indica, Japonica, Aus etc. and differed significantly on the basis of duration and genetic makeup. Thus, the genotype had strong Genetic*Environment (G*E) interaction that influenced their amylose and TS content. The glycemic index value is known to be influenced by genetic makeup and environmental factors [34] and storage conditions of the rice grains.
3.4 Enzyme assay:
The Starch synthase and Starch branching enzyme are known to be key enzymes of starch biosynthesis pathway in rice. The previous studies in Starch synthase have shown that the key activity of Starch synthase enzyme is correlated to amylopectin synthesis controlled by SSI, SSII, SSIII, and SSIV isoforms of SS gene [22]. Similarly, the activity of starch branching enzyme was known to be correlated with amylopectin synthesis controlled by BEI, BEIIa, BEIIb isoforms of BE gene [22].
Starch synthase: The activity of starch synthase enzyme was recorded for selected rice genotypes (Fig. 4). It is reported to be the value of 17.8 nmol/ (mg protein)/min in Oryza sativa cv. Fujihikari [25] while we found the highest value of starch synthase in Ashmber [9.385 nmol/ (mg protein)/min] and the lowest value in Makro [4.64 nmol/ (mg protein)/min]. we discovered a correlation between starch synthase activity in developing grains and GI. As varieties with high starch synthase are showing high GI values. The only exception observed in IR 64 as it has low GI but it was showing relatively high starch synthase activity.
Figure 4. Enzyme assay of starch synthase in selected rice genotypes
Branching enzyme: The branching enzyme assay was conducted in selected six genotypes (Fig 5). [25] reported value of 4540 nmol/ (mg protein)/min in Oryza sativa cv. Fujihikari. According to our research, Makro has the greatest branching enzyme value [2887.972 nmol/ (mg protein)/min] and Swarna has the minimum branching enzyme value [1928.852 nmol/ (mg protein)/min]. in our study branching enzyme activity in developing grain was found to correlate with GI value in Ashmber, Madhuraj 55 and Swarna.
Figure 5. Enzyme assay of branching enzyme in selected rice genotypes
3.5 Expression analysis of genes
The expression of nine genes- AGPL 1, AGPS 1, SS IIa, GBSS II, Wx, BE IIa, SUT 1, SUT 4 and SUT 5 were studied in flag leaf tissue of six diverse rice genotypes (Fig. 6 and Table 5)) and the results were correlated to the AC, GI and enzyme activity in developing grains. The expression of genes encoding ADP-glucose pyrophosphorylase large subunit or AGPL1 (regulatory subunit) and the catalytic component of the AGPase enzyme known as AGPS 1 (ADP-glucose pyrophosphorylase small subunit) was studied. These two genes together catalyse the process to transform ATP and glucose-1-phosphate into ADP-glucose and inorganic pyrophosphate (Ppi) in amyloplast [35, 36]. We found that these genes were expressed in all the six genotypes suggesting the occurrence of starch synthesis process in the flag leaf tissue. From this ADP glucose, leaf will produce starch, which will then be transported to sink organs (seed, endosperm) to form storage starch. So, it may be concluded that every genotype followed the similar initial stage for starch production. However, the variations for GI values and AC were present in these genotypes. It might suggest that the expression of these genes is not the cause of the variation in GI and AC levels.
The gene encoding SS IIa encodes starch synthase enzyme whose function is to elongate amylopectin short chains. SSIIa is known to be involved in amylopectin synthesis as loss of SSIIa in rice is reported to reduce amylopectin content [37]. SS IIa was expressed in all the genotypes except Swarna. There is another gene which is involved in amylopectin synthesis- BE IIa. BE IIa encodes Starch branching enzyme which produces amylopectin by forming branched glucans and starting an α-1,6-linkage through the transfer of liner glucan [21]. The BEIIa displayed variable expression among the rice genotypes with expression only in Ashmber, Makro, Noh hai and Swarna.
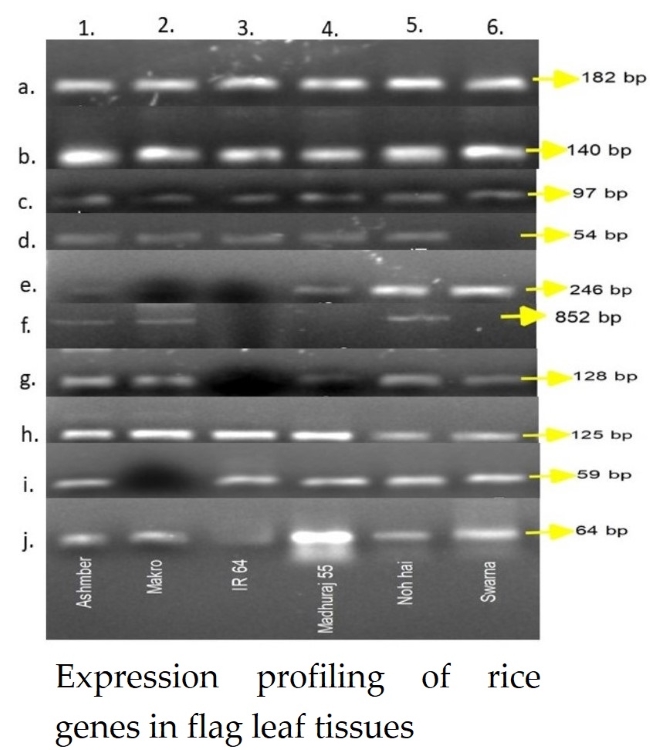
Figure 6: Expression profiling of 10 genes [a.- actin
(housekeeping gene), b.-AGPL1, c.-AGPS1, d.-SS2a, e.-GBSS2,
f.-Wx locus, g.-BE2a, h.-SUT1, i.-SUT4, j.-SUT5] in 6 rice genotypes
[1.-Ashmber, 2.-Makro, 3.-IR 64, 4.-Madhuraj 55, 5.-Noh hai,
6.-Swarna] in flag leaf tissue at 55 days after sowing.
Table 5. On the basis of the biochemical analysis and the phenotypic evaluation, six genotypes were selected for the enzyme assay and gene expression analysis.
S. No. | Genotype | Parentage | Biochemical analysis |
| Grain quality traits | |||||
AC | Total starch | GI |
| Grain length (mm) | Grain width (mm) | L/B ratio | Chalkiness | |||
1 | Ashmber | Indica | 23.17 | 59.64 | 88.44 |
| 6.5 | 2.2 | 2.95 | 7 |
2 | Makro | Indica | 24.88 | 70.3 | 67.37 | 7.9 | 2.8 | 2.82 | 5 | |
3 | IR 64 | Indica | 19.75 | 65.47 | 63.46 | 6.1 | 2.1 | 2.90 | 5 | |
4 | Madhuraj 55 | Indica | 25.18 | 64.37 | 62.79 | 7.4 | 2.4 | 3.08 | 7 | |
5 | Noh hai | Indica | 5.27 | 57.09 | 82.95 | 7.1 | 2.3 | 3.09 | 9 | |
6 | Swarna | Indica | 21.89 | 62.4 | 69.27 | 5.1 | 2.2 | 2.32 | 9 | |
GBSS II encodes granule-bound starch synthase enzyme II which is involved in amylose synthesis as it adds ADP glucose units in a linear chain by α-1,4-linkage [38]. GBSS II was only expressed in the Madhuraj55, Noh hai and Swarna, with relatively less expression in Madhuraj-55 (low GI genotype) as compared to the other two. Wx 6 gene encodes granule bound starch synthase I which primarily controls amylose synthesis in seed endosperm by generating linear glucan chain. GBSS I is also responsible for gel consistency (GC) regulation and amylose molecule size modulation [39, 40]. The expression of Wx was recorded in Ashmber, Makro and Noh hai varieties. The allelic variation waxy locus, with SNPs in Int 1-1 and EX 6-62 is known to influence AC of rice grain [41]. SUT 1 encodes Sucrose transporter 1, serves to retrieve sucrose from the phloem apoplasm along the transport pathway and loads sucrose synthesised in the scutellum into the sieve elements. SUT1 plays a role in rice plant germination, early seedling growth, and development [16]. All genotypes showed similar expression of SUT 1 gene. Only Noh Hai and Swarna displayed a lesser expression than the rest of the genotypes. SUT 4 encodes sucrose transporter 4, which is responsible for sucrose transport from the scutellum to the young developing shoots and roots as well as sucrose loading into the sheath phloem of the upper leaves during the post-heading period. SUT4 has a significant impact on plant development because its knockout effect reduces plant height and tiller number and causes yield loss [42]. It was expressed in all genotypes except Makro. SUT 5 encodes the sucrose transporter 5, which is crucial for delivering sucrose as a carbon source for growing tissues or possibly to supply sucrose to temporary storage tissues. It was expressed in all genotypes except IR 64 with higher level of expression in Madhuraj 55. SUT 5 gene knockout causes yield loss, increased endosperm chalk in the caryopsis, and decreased seed setting [43].
Similar gene expression was observed in Ashmber and Makro for SS 2a, BE 2a, SUT1 and SUT5 but the SUT4 gene did not express in Makro is the distinction. As found earlier, Makro has high AC and low GI, compared to Ashmber's high GI and AC. These variations solely differ in terms of GI value and SUT4 expression. SUT 4 loads sucrose into the upper leaves' sheath phloem during the post heading stage. Therefore, it is possible that poor loading of sucrose/starch onto endosperm led to dense deposition of amylose in grains of Makro.
A similar level of expression of SS 2a and SUT 4 genes was seen in Madhuraj 55 and Noh hai. However, they differ in how other genes are expressed; for example, in Madhuraj 55 high level of expression of SUT5, upregulated SUT1, low GBSS2, and no expression of BE2a or the waxy locus gene was observed. While Noh hai exhibits waxy locus gene expression, upregulated GBSS2 and BE2a expression, low SUT1 expression, and low SUT5 expression. As biochemical studies indicated that Noh hai has a low AC and high GI genotype, whereas Madhuraj-55 is a high AC and low GI variety. Low expression of the sucrose transporters SUT1 and SUT5 may be the reason for Noh hai's high GI value and low AC. It indicates that lesser amount of sucrose was transported to endosperm and whatever little sucrose is reaching endosperm it may be utilised for amylopectin synthesis rather than amylose.
Swarna and IR 64 are the mega varieties that expressed differential pattern in their gene expression and GI and AC values. The lesser expression of BE 2a and absence of waxy locus explains the intermediate AC and glycemic index values of these genotypes. These genotypes are known to have better milling percentage and head rice recovery also, which suggests that rice genotypes with less stickiness, intermediate AC, glycemic index and better head rice recovery (HRR) values are more preferred by farmers and rice millers.
4. Conclusions
The result of the present study suggests that the AC and TS values are not determined by the expression of a single gene but are affected by numerous other factors. In addition to maintaining the activity levels of individual enzymes, the gene–gene interactions might play an essential role in the metabolic regulation of starch biosynthesis. Amylose, amylopectin, and TS concentration were significantly influenced by sucrose transporters, which in turn affect the GI value. However, no relation was found in the enzyme activity of developing grains and gene expression flag leaf tissue indicating the stage and tissue specific expression of amylose and amylopectin synthesis related genes. The detailed study of the difference in the proteins of amylose and amylopectin synthesis and packaging in different rice varieties will facilitate the proper undertaking of the molecular factors determining glycaemic index value of rice grains.
Authors’ contributions
Conceptualization, A.K. and S.B.; Phenotyping biochemical estimation and expression profiling, A.M. and S.B.; Data analysis and software, T.B.; Validation and statistical analyses, V.P. and R.R.S.; Estimation of amylose content and glycemic index, A.M., P.R. and S.S.R.; Resources and lab facilities, S.V. and S.B.; Writing–Review and Editing of manuscript, A.M., S.B.; S.S.R. and V.P.; Visualization, S.B.; Supervision, S.B.V.; Project administration, S.B.; Provided support for standardization of methodology for GI estimation, U.M.S. and V.S.
Acknowledgements
We express our gratitude to the Richariya Laboratory, Department of Plant Molecular Biology and Biotechnology for creating a supportive environment that facilitated the laboratory analyses.
Funding
This research received no grant from any funding agency, it was partly funded by DBT PG teaching program by Dept. of Biotechnology, GOI, New Delhi
Availability of data and materials
All necessary data supporting the conclusions of this article will be available from the authors without undue reservation.
Conflicts of interest
The authors declare no conflict of interest exist.
References
1.
Curá, J.A.; Jansson, P.E; Krisman C.R.
Amylose is not strictly linear. Starch/Stärke, 1995, 47, 207-209.
http://dx.doi.org/10.1002/star.19950470602.
2.
Gallant, D.J.; Bouchet, B.; Baldwin,
P.M. Microscopy of starch: evidence of a new level of granule organization.
Carbohydr. Polym, 1997. 32 (3–4),177-191. https://doi.org/10.1016/S0144-8617(97)00008-8.
3.
Colleoni, C.; Dauvillée, D.; Mouille, G.
Genetic and biochemical evidence for the involvement of α-1,4
glucanotransferases in amylopectin synthesis. Plant Physiol. 1999, 120(4), 993-1004. https://doi.org/10.1104/pp.120.4.993.
4.
Gunning A.P.; Giardina T.P; Faulds,
C.B.; Juge, N.; Ring, S.G.; Williamson, G.; Morris, V.J. Surfactant-mediated
solubilisation of amylose and visualisation by atomic force microscopy.
Carbohydr. Polym. 2002. 51(2),177-182. https://doi.org/10.1016/S0144-8617(02)00126-1.
5.
Myers, A.M.; Morell, M.K.; James, M.G.;
Ball, S.G. Recent progress toward understanding biosynthesis of the amylopectin
crystal. Plant Physiol. 2000, 122 (4), 989–998. https://doi.org/10.1104/pp.122.4.989.
6.
Nakamura, Y. Towards a better
understanding of the metabolic system for amylopectin biosynthesis in plants:
rice endosperm as a model tissue. Plant Cell Physiol, 2002. 43 (7), 718–725. https://doi.org/10.1093/pcp/pcf091.
7.
Chen, Yi.; Mei, W.; Ouwerkerk, B.F.P.
Molecular and environmental factors determining grain quality in rice. Food
Energy Secur. 2012. 1 (2), 111-132. https://doi.org/10.1002/fes3.11.
8.
Lee, S.K.; Lee, J.; Jo, M.; Jeon, J.S.
Exploration of sugar and starch metabolic pathway crucial for pollen fertility
in rice. Int. J. Mol. Sci. 2022, 23(22). https://doi.org/10.3390/ijms232214091.
9.
Shimoyanagi, R.; Abo, M.; Shiotsu, F.
Higher temperatures during grain filling affect grain chalkiness and rice
nutrient contents. Agronomy, 2021, 1(7), 1360. https://doi.org/10.3390/agronomy11071360.
10.
Ahmed, N.; Tetlow, I.J.; Nawaz, S.;
Iqbal, A.; Mubin, M.; ul Rehman, M.S.N.; Butt, A.; Lightfoot, D.A.; Maekawa, M.
Effect of high temperature on grain filling period, amylose content and
activity of starch biosynthesis enzymes in endosperm of basmati rice. J. Sci. Food
Agric. 2015, 95(11), 2237–2243. https://doi.org/10.1002/jsfa.6941.
11.
Seung, D. Amylose in starch: towards an
understanding of biosynthesis, structure and function. New Phytologist. 2020. 228 (5),1490-1504. https://doi.org/10.1111/nph.16398.
12.
Fujita,
N.; Yoshida, M.; Asakura, N.; Ohdan, T.; Miyao, A.;
Hirochika, H.; Nakamura, Y. Function and characterization of starch
synthase I using mutants in rice. Plant Physiol. 2006. 140(3), 1070–1084. https://doi.org/ 10.1104/pp.105.071845.
13.
Jeon, J.S.; Ryoo, N.; Hahn, T.R.; Walia,
H.; Nakamura, Y. Starch biosynthesis in cereal endosperm. Plant Physiol. Biochem.
2010, 48(6), 383-392. https://doi.org/10.1016/j.plaphy.2010.03.006.
14.
Hu, Z.; Tang, Z.; Zhang, Y.; Niu, L.;
Yang, F.; Zhang, D.; Hu, Y. Rice SUT and SWEET Transporters. Int. J. Mol. Sci.
2021, 18,
22(20), 11198. https://doi.org/10.3390/ijms222011198.
15.
James, M.G.; Denyer, K.; Myers, A.M.
Starch synthesis in the cereal endosperm. Curr. Opin. Plant Biol. 2003. 6(3), 215-22. https://doi.org/10.1016/s1369-5266(03)00042-6.
16.
Scofield, G.N. The role of the sucrose
transporter, OsSUT1, in germination and early seedling growth and development
of rice plants. J. Exp. Bot, 2007. 58(3), 483–495. https://doi.org/10.1093/jxb/erl217.
17.
Pfister, B.; Zeeman, S.C. Formation of starch
in plant cells. Cell Mol Life Sci. 2016, 73(14),
2781–2807. https://doi.org/10.1007/s00018-016-2250-x.
18.
Naseer, B.; Naik, H.R.;
Hussain, S.Z.; Shikari A.B.; Noor, N. Variability in waxy (Wx) allele, in-vitro
starch digestibility, glycemic response and textural behaviour of popular
Northern Himalayan rice varieties. Sci. Rep. 2021, 11(1), 12047.
https://doi.org/10.1038/s41598-021-91537-0.
19.
Tetlow, I.J.; Beisel, K.G.; Cameron, S.;
Makhmoudova, A.; Liu, F.; Bresolin, N.S.; Emes, M.J. Analysis of protein complexes in wheat amyloplasts
reveals functional interactions among starch biosynthetic enzymes.
Plant Physiol. 2008, 146 (4), 1878–1891. https://doi.org/10.1104/pp.108.116244.
20.
Zhang, C.; Yang, Y.;
Chen, S.; Liu, X.; Zhu, J.; Zhou, L.; Lu, Y.; Li, Q.; Fan, X.; Tang, S.; Gu,
M.; Liu, Q. A rare Waxy allele coordinately improves rice eating and cooking
quality and grain transparency. J.
Integr. Plant Biol. 2021, 63(5), 889–901. https://doi.org/10.1111/jipb.13010.
21.
Tetlow, I. J.; Emes, M.
J. A review of starch‐branching enzymes and their role in amylopectin
biosynthesis. IUBMB life, 2014,
66(8), 546-558. https://doi.org/10.1002/iub.1297.
22.
Ohdan, T.; Francisco, P.B.; Sawada, T.; Hirose,
T.; Terao, T.; Satoh, H.; Nakamura, Y. Expression profiling of genes involved
in starch synthesis in sink and source organs of rice. J. Exp. Bot. 2005. 56 (422), 3229–3244. https://doi.org/10.1093/jxb/eri292.
23.
Juliano, B.; Perez, C.; Blakeney, A.;
Castillo, T.; Kongseree, N.; Laignelet, B.; Webb, B. International cooperative
testing on the AC of milled rice. Starch. 1981. 33(5), 157-162. https://doi.org/10.1002/star.19810330504.
24.
Goni, I.; Garcia-Alonso, A.;
Saura-Calixto, F.A starch hydrolysis procedure to estimate glycemic index.
Nutr. Res. 1997, 17(3), 427-437.
https://doi.org/10.1016/S0271-5317(97)00010-9.
25.
Nakamura, Y.; Kazuhiro. Y.; Shin Young,
P.; Toshihide, O. Carbohydrate metabolism in the developing endosperm of rice
grains. Plant Cell Physiol. 1989. 30 (6), 833–839. https://doi.org/10.1093/oxfordjournals.pcp.a077813.
26.
Rio, D.C.; Ares, M.; Hannon, G.J.;
Nilsen, T.W. Purification of RNA using TRIzol (TRI reagent). Cold Spring Harbor
Protocols, 2010, 6.
https://doi.org/10.1101/pdb.prot5439.
27.
Cheng, C.; Ali, A.; Shi, Q.; Zeng, Y.;
Tan, X.; Shang, Q.; Huang, S.; Xie, X.; Zeng, Y. Response of chalkiness in high-quality rice
(Oryza sativa L.) to temperature across different ecological regions. J. Cereal
Sci. 2019, 87, 39-45.
https://doi.org/10.1016/j.jcs.2019.02.009.
28.
Cruz, M.; Arbelaez, J.D.; Loaiza, K.;
Cuasquer, J.; Rosas, J.; Graterol, E. Genetic and phenotypic characterization
of rice grain quality traits to define research strategies for improving rice
milling, appearance, and cooking qualities in Latin America and the Caribbean.
Plant Genome, 2021, 14(3),
e20134. https://doi.org/10.1002/tpg2.20134.
29.
Lapis, J.R.; Cuevas,
R.P.O.; Sreenivasulu, N.; Molina, L. (). Measuring Head Rice Recovery in Rice.
In: Sreenivasulu, N. (eds) Rice Grain Quality. Methods in Molecular Biology,
vol 1892. Humana Press, New York, NY. 2019. https://doi.org/10.1007/978-1-4939-8914-0_5.
30.
Jinsong, B. "Rice
milling quality." Rice.
AACC International Press, 2019, 339-369.
31.
Sitoe, H.M.; Zhang, Y.; Chen, S.; Li,
Y.; Ali, M.; Sowadan, O.; Karikari, B.; Liu, E.; Dang, X.; Qian, H.; Hong D.
Detection of QTLs for plant height architecture traits in rice (Oryza sativa
L.) by association mapping and the RSTEP-LRT method. Plants. 2022, 11(7), 999. https://doi.org/10.3390/plants11070999.
32.
Xu, X.; Zhang, M.; Xu, Q.; Feng, Y.;
Yuan, X.; Yu, H.; Wang, Y.; Wei, X.; Yang, Y. Quantitative trait loci
identification and genetic diversity analysis of panicle structure and grain
shape in rice. Plant Growth Regulation, 2020. 90, 89–100.
https://doi.org/10.1007/s10725-019-00549-4.
33.
Atkinson, F.S. International tables of
glycemic index and glycemic load values 2021: a systematic review. Am. J. Clin.
Nutr, 2021, 114(5), 1625–1632. https://doi.org/10.1093/ajcn/nqab233.
34.
Beckles, D.; Thitisaksakul, M. Use of biotechnology
to engineer starch in cereals. Encyclopaedia Biotechnol. Agric. Food, 2014, 1, 10.
https://doi.org/10.1081/e-ebaf-120051354.
35.
Okita T.W. Is there an alternative
pathway for starch synthesis? Plant Physiol. 1992. 100, 560-564. https://doi.org/
10.1104/pp.100.2.560.
36.
Greene, T.W.; Hannah, L.C. Maize
endosperm ADP-glucose pyrophosphorylase SHRUNKEN2 and BTITTLE2 subunit
interactions. Plant Cell, 1998, 10(8),1295-1306. https://doi.org/10.2307/3870641.
37.
Miura, S.; Crofts, N.; Saito, Y.;
Hosaka, Y.; Oitome, N.F.; Watanabe, T.; Kumamaru, T.; Fujita, N. Starch
synthase IIa-deficient mutant rice line produces endosperm starch with lower
gelatinization temperature than japonica rice cultivars. Front. Plant Sci,
2018, 9. https://doi.org/10.3389/fpls.2018.00645.
38.
Hirose, T.; Terao, T. A comprehensive
expression analysis of the starch synthase gene family in rice (Oryza sativa
L.). Planta. 2004. 220(1), 9-16. https://doi.org/10.1007/s00425-004-1314-6.
39.
Biselli, C.;
Cavalluzzo, D.; Perrini, R.; Gianinetti, A.; Bagnaresi, P.; Urso, S.; Orasen,
G.; Desiderio, F.; Lupotto, E.; Cattivelli, L.; Valè, G. Improvement of
marker-based predictability of apparent amylose content in japonica rice
through GBSSI allele mining. Rice. 2014,
7(1), 1-18. https://doi.org/10.1186/1939-8433-7-1.
40.
Wanga, Z.; Luoa, Q.; Jianga, X.; Wub,
X.; Xua, H.; Zhua, S. Diversity analysis of the Waxy gene in Oryza sativa L.
“Guizhou HE”, Guizhou Rice Research Institute, Guizhou Academy of Agricultural
Sciences, Guizhou University China, 2021, Sci. Asia. 47, 434–440.
https://doi.org/10.2306/scienceasia1513-1874.2021.055.
41.
Zhang, C.; Zhu, J.; Chen, S.; Fan, X.;
Li, Q.; Lu, Y. Wxlv, the ancestral allele of rice Waxy gene. Proceedings. 2019,
36(1), 140. https://doi.org/10.3390/proceedings2019036140.
42.
Chung, P.; Hsiao, H.H.; Chen, H.J. Influence of temperature on the expression of
the rice sucrose transporter 4 gene, OsSUT4, in germinating embryos and
maturing pollen. Acta Physiol. Plant, 2014, 36,
217–229. https://doi.org/10.1007/s11738-013-1403-x.
43.
Aoki, N. The sucrose transporter gene
family in rice. Plant Cell Physiol, 2003, 44(3), 223–232.
https://doi.org/10.1093/pcp/pcg030.
This work is licensed under the
Creative Commons Attribution
4.0
License (CC BY-NC 4.0).
Abstract
The
Glycemic Index (GI) is an important quality trait of rice that determines
consumer preference. GI is dependent on the relative proportion of
amylose/amylopectin, soluble and resistant starch which in turn affects the
ease of hydrolysis of the starch. To understand the relation between amylose
content (AC) and GI the activity of starch synthase (SS) and branching enzyme
(BE) was studied in the developing grain at the milking stage along with the
expression analysis of 9 genes encoding enzymes involved in amylose synthesis,
amylopectin synthesis and sucrose transport in flag leaf tissue. Expression
analysis was done in 6 genotypes with high and low GI values viz., Ashmber-GI
88.4, Makro- GI 67.3, IR 64- GI 63.4, Madhuraj 55- GI 62.7, Noh hai- GI 82.9
and Swarna- GI 69.2. Out of the 6 genotypes the activity of SS enzyme, related to
amylopectin synthesis, was found highest in low GI rice genotypes (Makro,
Madhuraj 55 and Swarna) and highest in Ashmber and Noh hai (high GI genotypes).
The activity of BE was recorded maximum in Makro, Ashmber and IR 64 and minimum
in Swarna, establishing the role of SS enzymes in the formation of soluble
starch. The expression profile indicated
that the amylose and total starch (TS) content are not determined by a major
gene but are affected by the interaction of sucrose transporters and synthase
enzymes that in turn determined amylose, amylopectin, and TS concentration. The gene expression and enzymatic activity showed
interactions in the metabolic regulation of starch biosynthesis, transport, and
packaging grains, which significantly influenced the glycaemic index.
Abstract Keywords
Starch
biosynthesis, glycemic index, amylose content, sucrose transporter genes, rice.
This work is licensed under the
Creative Commons Attribution
4.0
License (CC BY-NC 4.0).
This work is licensed under the
Creative Commons Attribution 4.0
License.(CC BY-NC 4.0).



